In the first part “MACRAMÉ Data Shepherding – an Approach to the centralised Management of Research Information & Knowledge”, the principle data management concept of MACRAMÉ were outlined, including an elaboration of MACRAMÉ’s implementation of the FAIR principles for internal data sharing, and two new data management tools (i.e. (a) the MACRAMÉ Registry and (b) the instance map tool) as developed in collaboration with the BIO-SUSHY Project. The two tools were used in the first year of the project, to plan the workflows and study designs of the five Use-Cases and to document the status of the data, which was provided by the partners in their internal formats. Thus, these tools were focusing on the F (Findable) and A (Accessible) of FAIR in the form of a one-stop shop for all (Meta)Data (i.e. study designs, method description, protocols/standard operating procedures (SOP), data, presentations, and later publications and reports, as well as a visual representation of the links between different research outputs). To foster the I (Interoperable) and R (Re-usable), an additional third tool is now developed and introduced providing a programming library and scripts to automate workflows for translating (meta)data as provided by the partners into computer-actionable files. These files are structured into a still-to-be-completed harmonised MACRAMÉ (meta)data schema designed to fulfil requirements from data producers, as well as data users.
FAIR data sharing, independent of it being applied internally within a project or an organisation or for public sharing, requires harmonisation of data to achieve interoperability. But current lab workflows are not and, in some cases, probably never will be developed around standardised data schemas and exchange formats. Instead, they use customised files or electronic lab notebooks (ELNs) / laboratory information management systems (LIMS) with customised, partner-specific data models. MACRAMÉ is no different in this, with partners like EMPA and MyBiotech using very sophisticated and highly detailed but incompatible data management systems. To make data interoperable, previous projects used templates, in which each partner had to provide their data to the project, requiring an often manual translation of the partner-internal data-format to the project’s (meta)data specification and format. We do not want to derogate the importance of data reporting templates proposed for (nano)materials in the past (Exner et al., 2023 and references therein) in providing required information for nanosafety evaluation and regulation. However, agreements on these templates are very hard to achieve within a single project and almost impossible across projects, while the need to be applicable to different assays, including new developments, resulted in the templates often covering the bare minimum, only, while lacking information to fully evaluate the results by others. Additionally, the added burden is, in practice, often avoided or postponed, thus limiting the amount of FAIR data, and ultimately rendering any benefit from FAIRification during the project impossible.
To circumvent extra burden, we need to transition to an on-the-fly FAIRification approach, where harmonisation and machine-readability/actionability is achieved as part of the normal data collection process (see Exner et al., 2023). To enforce substantial changes of the internal data management systems, in order to comply with one common standard is not feasible and – even more importantly – not sensible, since it would require the retraining of personnel (potentially for each individual project) and would remove the flexibility from the laboratory workflows to report all parameters for internal management (e.g. internal IDs for chemicals, operator information, references to internal documents), without making the data model unmanageably complex and exposing confidential information. Instead, one could argue that harmonisation and interoperability require – in principle – simply the digital form of the (meta)data reporting, but not its specific structure. The FAIR principle accordingly requires that the data is reported according to domain-agreed community standards, only. As these standards are required to be well documented, translating from one to another is possible, similarly to translating between languages.
The FAIR principle accordingly requires that the data is reported according to domain-agreed community standards, only.
MACRAMÉ is building on this idea and accepts customised, partner-specific files as the primary data source. To still achieve harmonisation and interoperability, these files have – in most cases – still to be improved or complemented to:
- guarantee that all required metadata is available to make the data understandable for others, and
- that the data structure in the (meta)data file can be mapped to a standardised (meta)data schema.
How to achieve this improvement of the overall metadata coverage and completeness and create harmonised data from all the different resources is described here.
(Meta)Data Completeness
(Meta)data completeness in MACRAMÉ is primarily defined by the data requirements of the project-internal data users of the work package focussed on ‘Definition, Demonstration & Validation of MACRAMÉ Methods in Use-Cases’, who are performing safety and sustainability evaluations of the products of the five market-relevant MACRAMÉ Use-Cases. However, for final public sharing of the data without additional manual effort (i.e. on-the-fly FAIRification), cross-discipline and community specific (meta)data requirements (minimal reporting checklists and standards) have also to be considered from the start to achieve data completeness for internal and external data reuse. The first is achieved by discussing data availability and data needs in the Use-Case teams (including integration of learnings from other Use-Cases) supported by the instance maps visualising material and data flows. The latter is then added by consulting metadata standards endorsed by the FAIR community and relevant scientific communities and selection of the most appropriate ones guided by the data shepherds. For example, bibliographic and data provenance metadata requirements can be taken and adapted from different web standards and data services (e.g. GBIF, DataCite, schema.org and bioschemas.org, DDI). Again, it is important to note that the information requested by the standards needs to be provided but not necessarily in the exact same format or even data structure as used in the services. Even multiple standards can be combined (e.g. if they cover different aspects or are missing some but not the same metadata fields considered essential by MACRAMÉ). Additionally, standards can also be reused in parts if only substructures of the complete data schema are relevant (e.g. the fields related to authors, origin and supporting citation of the DataCite schema but not necessarily funding). Completeness with respect to all the data usages in MACRAMÉ is being established in parallel with the execution of the Use-Cases.
An important step was made during the 3rd MACRAMÉ Project Meeting in St. Gallen (11.-14. December 2023) and especially in a hands-on session lead by the work package on ‘MACRAMÉ Information & (Meta)Data Harmonisation, Processing & Sharing’: It looked at ways to define the products with the embedded advanced materials chemically and structurally as precisely as possible. Data exchange needs were first discussed for separate Use-Cases to identify important sources (i.e. project partners vs. public services, experimental vs. computational). The results of this metadata scoping exercise will now be combined into one central and global data model so that data users can check, if a specific (meta)data field is already planned to be collected or should be taken over from one to another Use-Case, and by whom it would be provided, using which method. Parts of an early version of this harmonised MACRAMÉ data schema is shown in Figure 1, which will be transformed into a JSON file during the process described below.
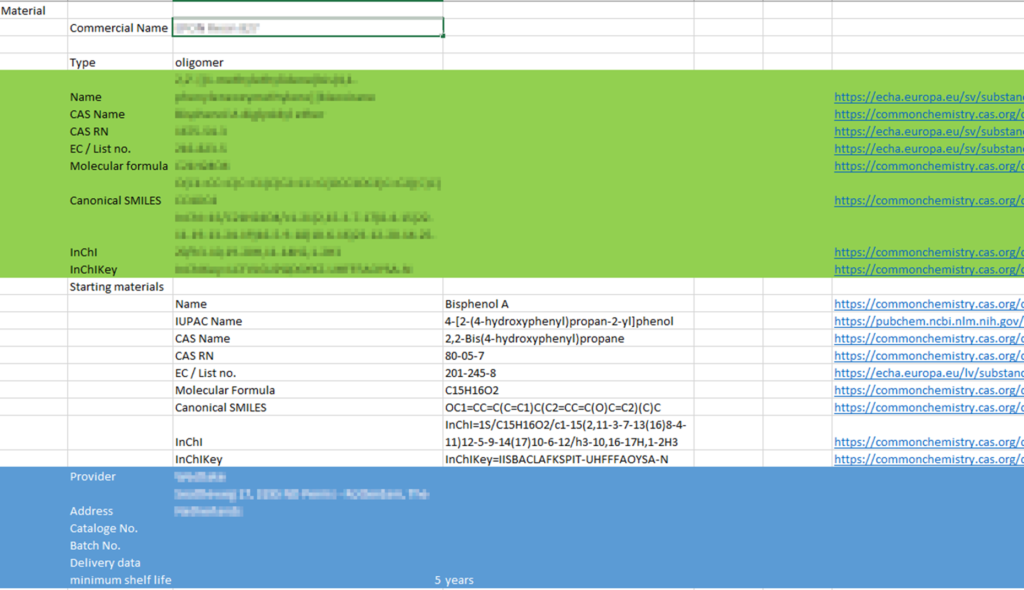
JSON has the advantage that the structure can easily be amended and locally re-structured to integrate new requirements that show up during further progression of the Use-Cases. Additionally, the data model documentation can be further enriched by including information on the expected data type, relationships between data entries, and even annotation with ontology terms essential for integrating MACRAMÉ data into the semantic framework of the materials data ecosystem. Combined with the automated procedures described in the next subsection, which is used for extracting data from partner-specific files, this setup provided the flexibility to generate the first versions of the datasets now and can be used to update these whenever improvements in data completeness and/or data documentation/annotation are requested by the data providers or users.
Data Collector
To achieve interoperability within MACRAMÉ, all (meta)data coming from the partners has now to be mapped onto the harmonised data schema and transformed accordingly. To reduce the time for doing this, the data collector software library is being developed by the Project’s data shepherd 7P9. This library is used to extract data from partner-specific data files, to automatically translate the data according to user-provided rules and to enrich the data by querying public services. We will demonstrate this with the Excel file used as starting point for the discussions in St. Gallen (see Figure 1). After the assumed data producer has uploaded a data file to the Google Shared Drive, currently used for knowledge exchange within MACRAMÉ, the semi-automated workflow visualised in Figure 2 is started.
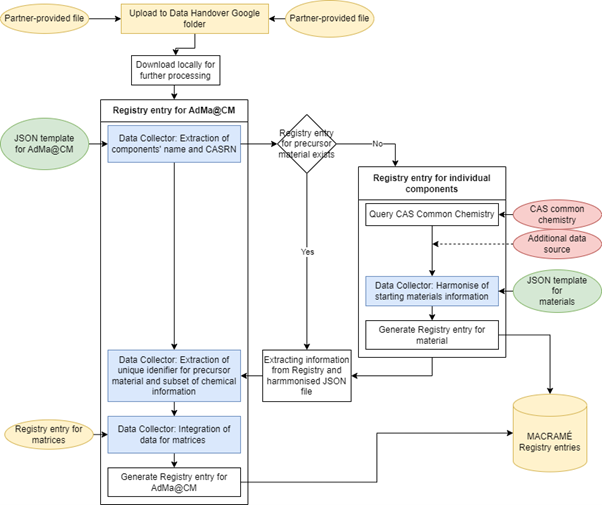
A customised script (in the F# language, see Figure 3) based on the data collector is used to fill the harmonised MACRAMÉ data schema with all information available in the file by accessing individual cells or blocks of cells in the spreadsheets. At the same time, the script performs basic checks on the content, like expected data type, to guarantee the consistency of the integrated data. In principle, the data collector could also directly access files, such as EMPA’s ELN, without the need for Excel files (or in any other format) for data exchange. However, this would need establishing a secure exchange between the two systems and exchange policies, which is out of scope of the MACRAMÉ Project.
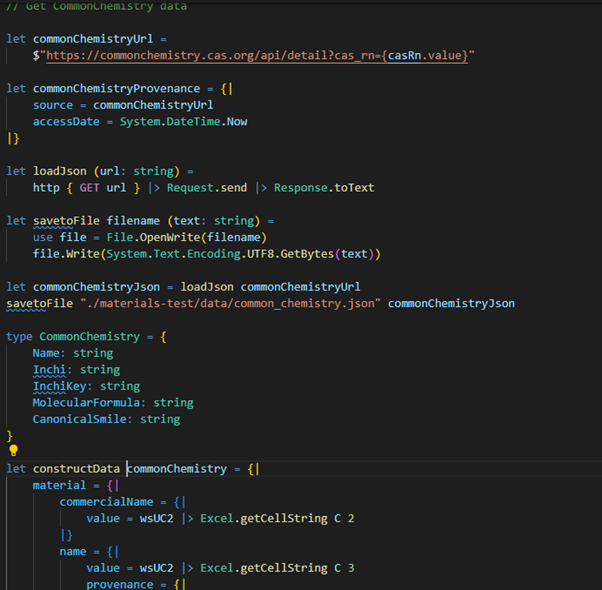
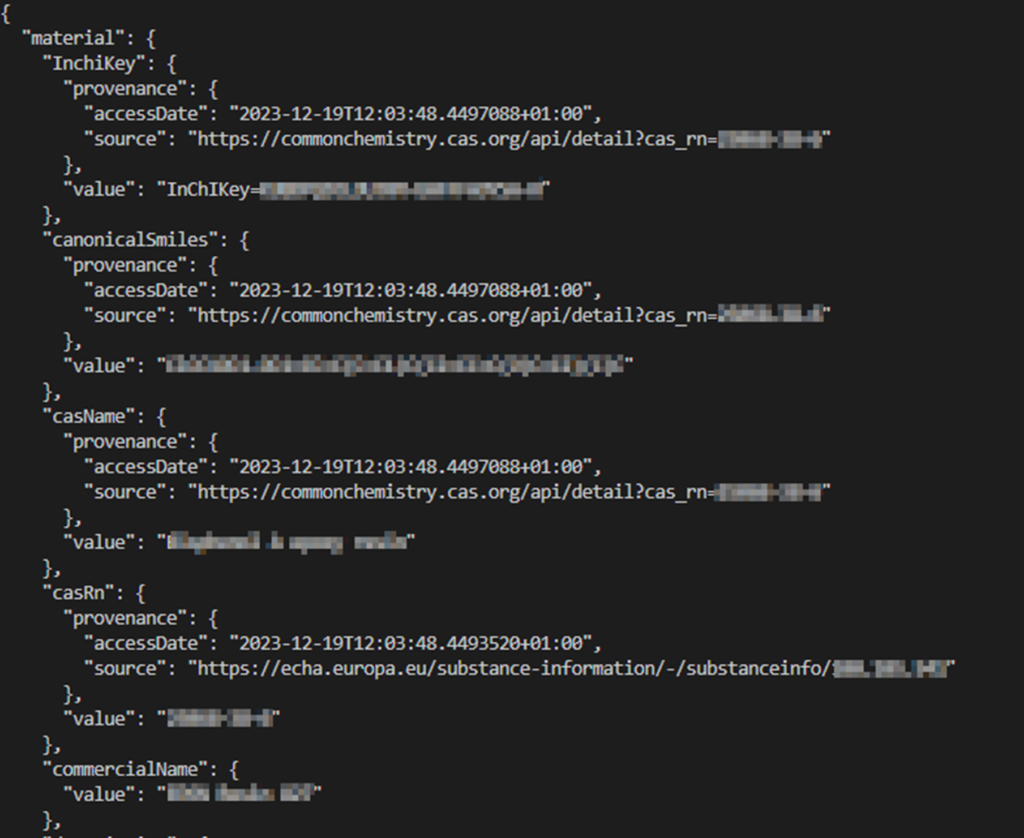
Figure 4 shows the resulting very simple JSON file. This is just for demonstration and the MACRAMÉ data schema is much more complex to cover all aspects of the AdMas and their product and biological matrices. Additional information, needed for complete documentation and (meta)data completeness not provided in the data files, would now have to be provided (e.g. in additional files of all kinds of formats).
In some cases, information on precursor materials is provided, a parallel workflow is started for each of them (see Figure 2). This extracts, if not already available from previous executions of the workflow, additional chemical identifiers (InChI, InChI key, SMILES) and basic physicochemical properties from web services like the Chemical Abstract Services Common Chemistry, PubChem and ECHA.
Please note that MACRAMÉ is encouraging or will later even enforce provision of provenance metadata to have the full trail to where the data came from and how it was processed. Unique identifiers assigned by the Registry and a subset of the precursor material information are then also stored as part of the AdMa data. In this way, the basic information on the material production including starting material identification is defined in the AdMa data file and additional data is accessible by following the link to the precursor material Registry entries. This characterisation data can then be complemented by further physicochemical characteristics as well as safety and life cycle assessment data generated in MACRAMÉ. After all this information is collected, it can then be accessed as is (e.g. as input for read-across and grouping) or translated once again into FAIR data formats used in public databased for long term storage or into standard reporting formats used in research and regulatory setting like the templates mentioned above or the OECD harmonised templates.
Concluding Remarks
As a conclusion of this second part of the MACRAMÉ Data Shepherding posts, we want to highlight a critical feature of the automatic procedure that profits from combining the Registry, instance maps and data collector. As described above, the harmonised MACRAMÉ (meta)data schema is not completely defined yet, since data requirements from safety and sustainability assessment will be refined alongside the progress in the Use-Cases. This is not only related to additional data needs but also to metadata completeness to fully document the production and testing to allow data reuse by other partners and finally open sharing. This additional (meta)data could be directly added to the Registry entries and the linked data files. However, the improvement would then apply only to these entries, while data providers would still generate incomplete (meta)data for new, related AdMas, which would then need to be amended later. Instead, updating the partner-specific files to integrate the additional information is improving the data management of the partners, and new data is then already complying with the higher quality standards. (Re)running the automatic harmonisation workflow integrates all data into new versions of the harmonised files of existing Registry entries or the first versions for new AdMa candidates. In that way, data harmonisation and on-the-fly FAIRification can be implemented and continuously enhanced with only minor modifications in the established experimental workflows at the partner institutions.
References:
Exner, T. E. et al. Metadata stewardship in nanosafety research: learning from the past, preparing for an “on-the-fly” FAIR future. Frontiers in Physics 11, (2023).






